Help
You can start building your network in several ways: Go to network page and input gene symbol in the input_field on the top right field of this page and press "Find":
After that, your gene/ncRNA node will appear on the canvas
If you cntrl+click (cmd+click for mac users) on this node, a table with upstream and downstream genes/ncRNA will appear in the right sidebar:
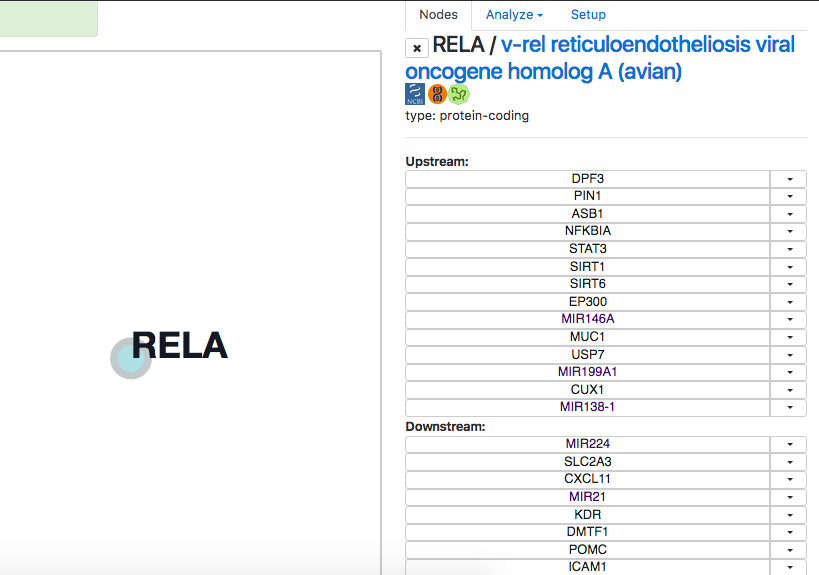
You can click on the upstream or downstream node, and it interactively appears on the canvas and also will contact other nodes, if there are such links.
Nodes shapes legend:
circle — gene/protein noderombus — ncRNA
squere — metablite
cross — drug
The second method is similar to the first option, but if you want input several genes/ncRNAs, -- just click "Upload" button:
Also, you can build a network according to the initial criterion. As a criterion can be: gene, disease, pathogenetic process. To select a criterion, you can go to the main page, click "Search" and than select the entities you are interested in:

Also you can build more complex criteria in this page. Using and combine many filters you can explore in details complex interconnections in cancer.
Analyze tab
Any network can be analyzed. To do this, there is a "Analyze" tab in the right side-bar:
1) Pathogenic process (PP) - molecular, cellular or tissue malfunction determining initiation, development and outcome of disease. Pathogenetic processes derived from Hallmarks of Cancer (HoC).
Processes are represented as an interactive table, which shows: the name, the number of links on this network, the number of interactions that enhance or weaken this process:
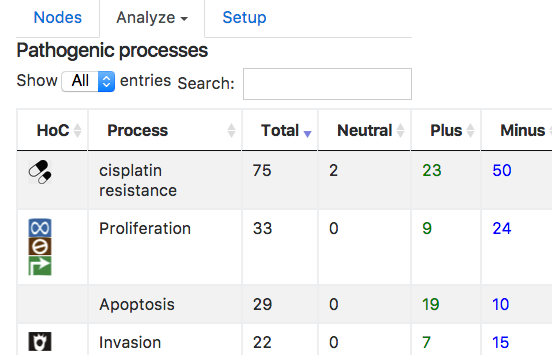
Biomarkers:
Biomarkers can be of three types: diagnostic, prognostic and drug resistance.
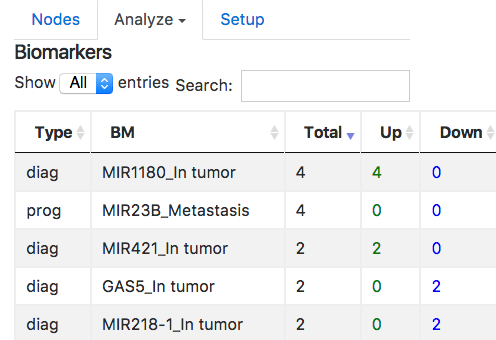
Diagnostic biomarker is used to identify whether a patient has a specific disease condition.
Prognostic biomarkers defined prognosis For example miR-21 upregulated and associated with invasion, migration and proliferation, in articles described correlation between OS (overall survival), PFS, poor prognosis
Predictive biomarkers defined treatments efficacy
Interactions associated with pathogenic processes and cancer hallmarks can determine drug efficacy. For instance, pathogenic processes, such as stemness and hypoxic response can determine chemoresistance, antiapoptotic microRNA activate array signaling pathways, including PI3K, AKT, JAK/STAT, mTOR and interfere with drug actions.
We collected associations in GRN (gene regulatory networks) context and pathogenic process related to drug efficacy.
Expression levels:
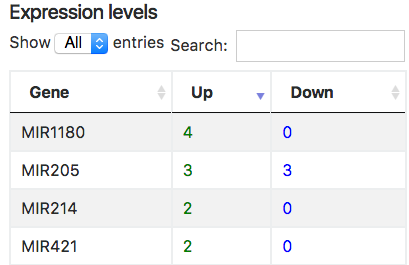
In ONCOIO each left node in interaction (if possible) has expression level (up or down) extracted from articles, for example

If
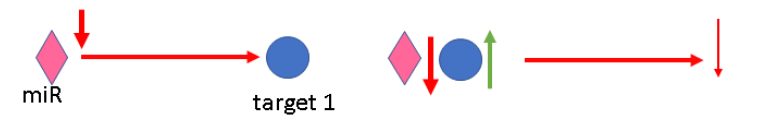
edge (interaction) is inhibited because left interactor was down-regulated or
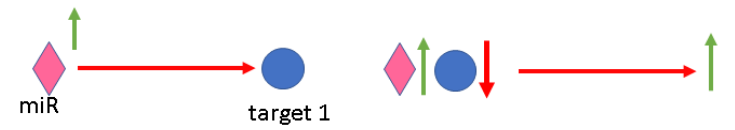
edge (interaction) is activated because left interactor was up-regulated
Also each edge associated with pathogenic process.
As we can show on the real examples: For interaction miR-142-3p => HMGB1 (PMID 27829233) “the expression of miR- 142 was obviously down-regulated in human cervical cancer tissues and a panel of cell lines.” and “miR-142 affects cervical cancer cell proliferation and invasiveness, and enhances cell apoptosis via directly targeting the expression of HMGB1” In ONCOIO downregulation miR-142-3p linked to disease (cervical cancer), processes (proliferation, invasion and apoptosis) with corresponding effect:
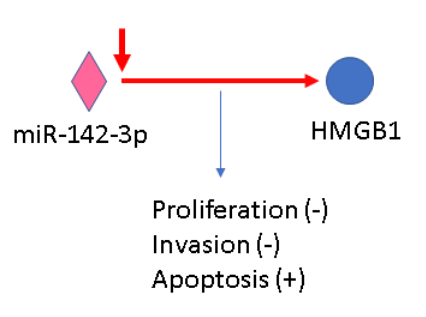
In this case miR-142-3p can not inhibit proliferation and invasion because its expression was down-regulated.
The pathogenic processes corresponding only to normal miR-142-3p function
Diseases:
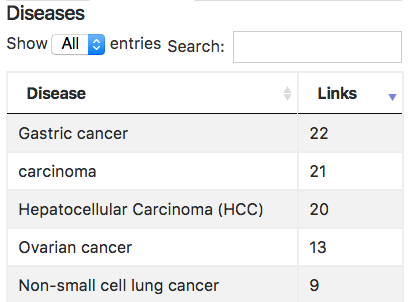
Visualize tab
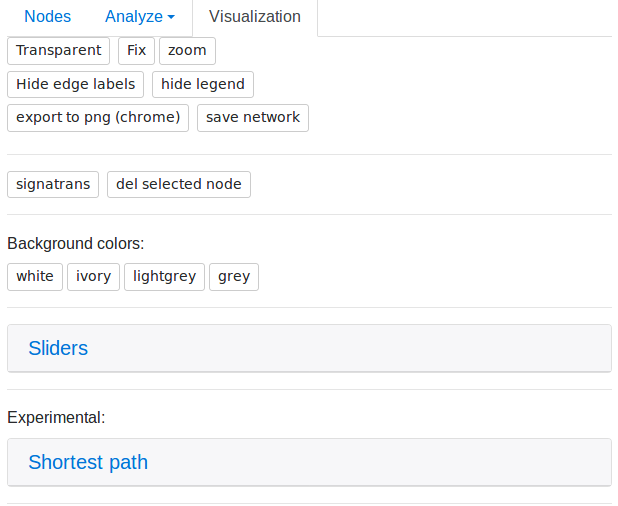
"Transparent"/"No transparent" button shows network edges more or less contrast
"Fix"/"Unfix" button fix (pin) or unfix network nodes
"Zoom button helps to find the network center
"Hide edge labels"/"Show edge labels" button hide or shows edge labels
"Hide legend"/"Show legend" button hide or shows legend
"Export to png (chrome)" button export network to large image in PNG format
"Save network" button save network in database by new name. This network can be used after and saved in user profile. All such networks have a permanent link.
"Sliders" button show sliders for change network topology.